Global environments are highly diverse and dynamic, offering many changes and adaptive challenges to creatures. However, DNA sequence variability engineered into plant and animal genomes is one design feature (of many) that allows creatures to rapidly deploy adaptive traits in response to sensing a broad range of environmental conditions.
Global environments are highly diverse and dynamic, offering many changes and adaptive challenges to creatures. However, DNA sequence variability engineered into plant and animal genomes is one design feature (of many) that allows creatures to rapidly deploy adaptive traits in response to sensing a broad range of environmental conditions.
An Introduction to Genetic Diversification
One of the earliest scientists to make big strides in recognizing and understanding DNA variability was Gregor Mendel (1822–1884), an Austrian monk whose work inspired early twentieth-century studies in genetics. He observed certain flower and seed trait variations in the offspring of peas and other garden plants following his controlled matings.1 In some cases, a progressive continuum of flower color could be noticed in the offspring. In other cases, the traits segregated in seemingly predictable statistical outcomes. We now know these to be simple or qualitative traits controlled by a single gene—a very rare occurrence in nature.
These qualitative traits in pea plant flower color were due to a changed pigmentation-related gene that produced white flowers (no pigment) rather than pigmented purple flowers.2 Because pea plants, like most animals, have two sets of chromosomes (one from the mother and the other from the father), even if a pea plant had only one good copy of a pigmentation gene, it could still produce purple flowers. If a plant had two copies of the altered gene, white flowers would result.

The key thing about Mendel’s work was that it demonstrated that genes or other chromosomal features could be shuffled around via matings in a population to create a diverse amount of variability. We also now know that nearly all traits in creatures, especially those that facilitate adaptation, are actually controlled by many genes and regulatory elements located across the genome at specific sites called loci (singular, locus). Genetic differences between loci variants are called alleles.
The potential for trait variability can be quite large due to the many possible allele combinations contributing to a trait. These types of traits are called polygenic (controlled by many genes) or quantitative traits (as compared to qualitative). These polygenic loci or alleles are typically referred to as quantitative trait loci (QTL) and are all interconnected in complex gene regulatory networks. Each locus contributes to different levels of variability in the trait being studied.
Quantitative traits and QTLs can be characterized in two different ways. The original approach was to perform controlled matings of a plant or animal to create a large population of offspring. Each individual was then characterized for visible traits (phenotypes) and had its genome mapped with DNA markers. Traits associated with a specific set of alleles (marker-connected loci) are determined by statistically analyzing the percent variability they contribute to a certain trait.
As genome sequencing became more common and efficient, this mapping strategy was largely replaced and/or supplemented by sequencing the entire genome of a large sample (cohort) of individuals from a population. Traits of interest are then correlated with variations in genome sequence in what are called genome-wide association studies. In fact, it is now possible to do this with populations from multiple environments. The outcome of such studies reveals both the location and DNA sequence of alleles connected to a variety of traits in organisms inhabiting different environments.
Creature Diversification
One reason why studying and understanding allele diversity in the genome is important is because it contributes to creature diversification, known by conventional scientists as speciation or adaptive radiation. The genetic foundation for this is realized by extrapolating allele diversity found in a cohort of individuals to an entire population of a creature where the diversity is harbored. Population-level genetic diversity is typically called standing genetic variation (SGV). Diversification occurs when a subsection of a population breaks off and pioneers a new environmental niche, carrying with it a subsection of the SGV of the founder population. If the breakaway group is small, that can eventually lead to inbreeding, which is detrimental to many types of higher animals but is less so to most plants.
A population shift can also occur if the environment that the founding population lives in is altered in some way. This can lead to a change in genome structures that are better adapted to the new conditions. However, contrary to standard Darwinian predictions, this often does not mean that adaptive variation favorable to the previous environment is lost. It merely isn’t predominant in the SGV, so a shift back to the prior condition is still possible. This kind of diversification has been demonstrated by a number of studies, several of which will be summarized in the following sections.
Examples of Standing Genetic Variation (SGV) in Adaptation
Cichlids
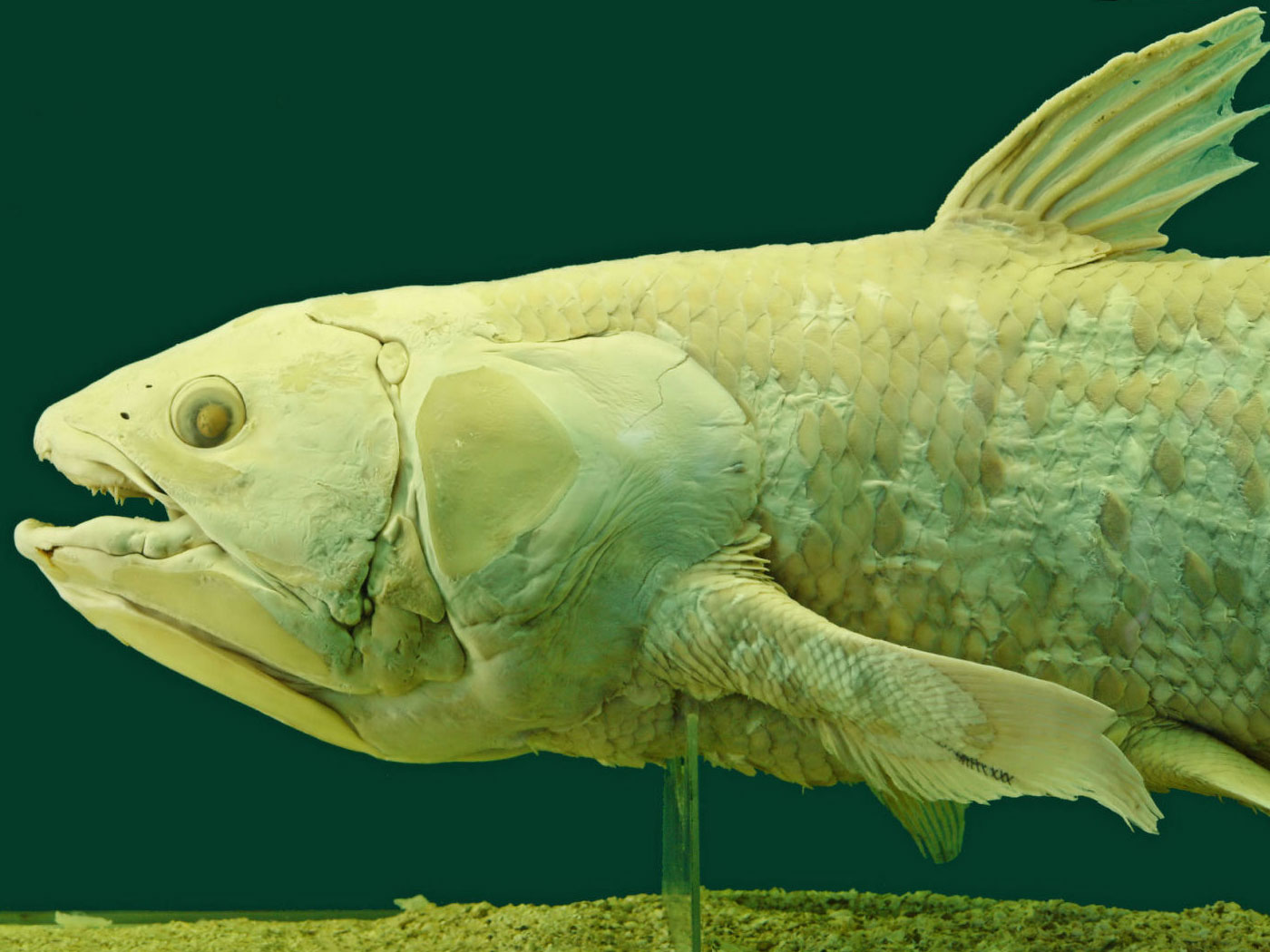
A stunning array of cichlid fishes have diversified in African lakes. The three largest lakes (Malawi, Tanganyika, and Victoria) contain cichlids that express well-adapted traits to eating plankton, algae, snails, insects, and even other fish. They also have distinct body shapes and coloration.

Neo-Darwinism speculates each of these cichlid forms emerged through slowly accumulated random genetic errors that allegedly supported a new beneficial trait that was selected for by the mystical agent of natural selection. However, recent studies have shown that the actual source of this adaptive radiation is preexisting, built-in genetic variation. In fact, the authors of a recent paper on cichlid adaptation wrote, “In summary, we identified genes with divergent alleles that originated before the adaptive radiation of Lake Victoria cichlids, suggesting the importance of SGV in their ancestral radiation of East African cichlids, and not just Lake Victoria cichlids.”3 In other words, the SGV that allowed for the adaptive radiation was ancestral and preexisting.
Not only does the SGV needed for adaptation preexist, or rather is pre-engineered and originally placed in creatures at creation by the Creator, but this process of diversification is strategically and repeatably deployed based on the environment that the cichlid is pioneering. In other words, cichlids adapted to similar environments separated by large amounts of geographic distance and time will deploy similar allelic configurations. This illustrates the repeatable nature of this process. The researchers stated, “It is clear that allelic polymorphisms are shared among different species of Lake Victoria cichlids due to recent adaptive radiation from the ancestral population with SGV,” and “the sharing of allelic polymorphisms is also observed across East African lakes and rivers.”3 In fact, the researchers go on to more clearly tell us what types of alleles are actually involved in specific modes of adaptation, saying, “The allelic diversity of genes related to sensory perception, including vision and olfaction [the sense of smell], is crucial for rapid speciation, because perception directly contributes to reproductive communication.”3
Marine Midges
Marine midges belong to the genus Pontomyia. They are flightless insects that inhabit marine coastal habitats and specifically thrive in the intertidal zone, salt marshes, and tidal pools along ocean shores. Not only are midges unique in that they are specialized insects that develop in saltwater, but their inability to fly, extreme sexual dimorphism, and very short adult lifespans of just a few hours make these creatures extremely unusual among insects in general. They are also somewhat ubiquitous, being found in the coasts of the Indian, Atlantic, and Pacific Oceans.

A recent study analyzed ecotypes of marine midges that have adapted themselves to a North Atlantic European habitat.4 At extreme northern latitudes, these midges differ from others in their egg-laying behavior and reproductive timing, involving both circadian and circalunar clocks. Circalunar clocks are innately engineered biological time-keeping mechanisms that allow organisms to monitor lunar phases and adjust adaptive mechanisms accordingly. In addition to the circalunar clock aspect, these creatures require their habitat to be exposed by the tide for successful egg laying. So, they live in intertidal zones along the European Atlantic coasts, which are dynamic and fluctuating environments and are maximally exposed during the low waters of spring tide days around the full moon and new moon. The emergence of adults is restricted to these circumstances by the creature’s circalunar clock, which tightly regulates the processes of development and maturation. This circalunar system also interfaces with an internal circadian clock based on the 24-hour solar cycle. This ensures emergence during only one of the two daily low tides. The adults reproduce immediately after emergence and die just a few hours later.
In this study, the researchers used a technique mentioned previously called QTL mapping. They genetically studied a population derived from a cross between lunar-sensitive and lunar-insensitive strains. As would be expected based on the fact that adaptive traits are complex, the researchers state, “We can conclude that lunar rhythmicity is not controlled by a single large-effect locus, but rather has an oligogenic [multiple genes with large effect] or possibly even polygenic basis [many genes].”4 They then did even more genetic testing through genome sequencing in the test population. They concluded that “ecotype formation is based on a complex polygenic architecture” and “adaptation in the North involves a re-assortment of existing standing genetic variation.”4 So, in the final analysis, this complex form of adaptation is based on many different genes and is also rooted in the exploitation of pre-engineered SGV.
Parrotbill Songbirds

Parrotbills are small, social birds that live in groups. Although they are found across the world, they are especially prevalent in diverse species/ecotypes across Southeast Asia, China, and Russia. In 2019, a Taiwanese group of researchers studied the sequenced genomes of 80 individuals involved in adaptation at differing latitudes in mainland China and Taiwan. It is noteworthy that the authors state at the beginning of the paper: “What kind of genetic variation contributes the most to adaptation is a fundamental question in evolutionary biology.”5 In their study, they found significant variability in many different groups of genes involved in the oxygen transport cascade,6 angiogenesis,7 respiratory organ modifications, blood hemoglobin content, metabolic processes related to thermoregulation, lipid metabolism, and ATP hydrolysis.8 Interestingly, they also detected diversity in biomolecules related to epigenetic modifications of chromatin (DNA complexed with proteins and RNA) associated with adaptation.

These results show that these birds are able to cope with environmental changes quickly by exploiting SGV across a wide range of genes and genomic regions for a broad diversity of specified physiological and morphological traits. Not only that, but these data also show that creatures with high levels of genetic diversity can persist in changing environments—providing an ongoing genomic resource to pioneer and colonize other novel environments. The significant role of SGV in adaptation in this parrotbill study has important implications for our understanding of adaptation in songbirds and creatures in general.
Sicilian Daisies
The Sicilian daisy is a highly adaptable wild plant that lives at multiple elevations in mountainous areas in Sicily, Italy. In a recent study, researchers wanted to assess the change in overall plant adaptation for this particular organism by relocating it to elevations outside its natural habitat on Mount Etna.9 They began by making crosses of plants growing in its natural habitat of 500 to 1,500 meters of elevation to get a good mix of the population’s SGV. Then they planted the offspring at an elevation of 2,000 meters.

They showed that significant amounts of hidden genetic variation in the founding population was utilized in adapting to the much higher elevation. Interestingly, a wide number of genes tripled in their levels of gene expression at the higher elevation compared to the beginning lower elevation. Many of these genes pertain to biosynthesis of important organic compounds related to plant adaptation along with genes connected to photosystems.
Significantly, the researchers gave the credit for this adaptation to preexisting (built-in) genetic variability as opposed to the evolutionary myth of random mutations. They said, “These results suggest that genetic variation important for rapid adaptation already segregates in the population, which makes evolutionary rescue [survival under environmental stresses] more likely than if adaptation were to rely on new mutation.”9 They go on to say that “these results suggest that standing genetic variation in plasticity [trait adaptability] could help populations to persist and adapt to novel environments, despite remaining hidden in native environments.”9
Summary
SGV is one of many divinely engineered features in living systems that allow diverse creatures to adapt themselves to a wide variety of environmental challenges and also enable them to pioneer new niches. Remarkably, much of this genetic variation is repeatable in its genomic configuration (according to the environment) and can remain hidden while organisms reside in a highly favorable environment. This hidden SGV can then be strategically deployed when stresses or new challenges are encountered. While the mechanisms utilized to both maintain and deploy SGV remain to be better elucidated, one thing we can say for sure is that the Creator, the Lord Jesus Christ, engineered and provided this variability for living systems and placed it in them at their creation about 6,000 years ago.
References
- Mendel, G. 1965. Experiments in Plant Hybridisation. Edinburgh and London: Oliver & Boyd.
- Hellens, R.P. et al. 2010. Identification of Mendel’s White Flower Character. PLoS One. 5 (10).
- Nakamura, H., M. Aibara, and M. Nikaido. 2023. Ancient Standing Genetic Variation Facilitated the Adaptive Radiation of Lake Victoria Cichlids. Genes and Genetic Systems. 98 (2): 93–99.
- Fuhrmann, N., C. Prakash, T. S. Kaiser. 2023. Polygenic Adaptation from Standing Genetic Variation Allows Rapid Ecotype Formation. eLife. 12: e82824.
- Lai, Y. T. et al. 2019. Standing Genetic Variation as the Predominant Source for Adaptation of a Songbird. Proceedings of the National Academy of Sciences USA. 116 (6): 2152–2157.
- An oxygen transport cascade is the physiological stepwise process that brings atmospheric oxygen into the body for cellular metabolism.
- Angiogenesis is the physiological process through which new blood vessels form from preexisting ones.
- ATP hydrolysis is the catabolic process where adenosine triphosphate (ATP) reacts with water to release energy, yielding adenosine diphosphate (ADP) and an inorganic phosphate. This releases a large amount of cellular energy to drive essential cellular activity, cellular transport, muscle contraction, and biosynthesis of key molecules.
- Walter, G. M. et al. 2023. Hidden Genetic Variation in Plasticity Provides the Potential for Rapid Adaptation to Novel Environments. New Phytologist. 239 (1): 374–387.
Dr. Tomkins is a research scientist at the Institute for Creation Research and earned his Ph.D. in genetics from Clemson University.