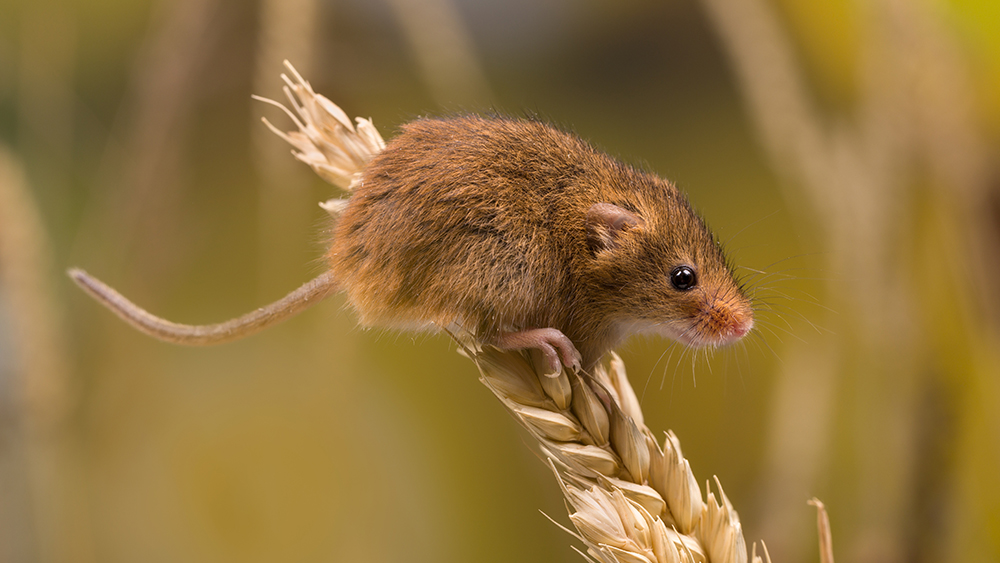
Darwin proposed that evolution happens externally, that the environment shapes organisms. But a growing amount of evidence suggests the opposite: Most changes happen because the organisms themselves sense, and react to, the environment. Thus, adaptation occurs internally because of superior design, not externally as a result of natural selection. A new report on gene regulation in mice intestines adds to the evidence of internal adaptation and design.1
Most changes happen because the organisms themselves sense, and react to, the environment. ![]()
Previous research on mice intestine cells established that, as these cells absorb and process different nutrients from the gut, they rapidly express various metabolic proteins in order to accurately match specific nutrients. If this observation was analyzed from a design standpoint, it would suggest that there should be an innate cellular system to link the detection of specific available nutrients in the gut lumen to some type of logic mechanism regulating the cell’s genes to produce specific proteins.
New research led by Dr. Patrick Varga-Weisz of the Babraham Institute at Cambridge demonstrated that this system does exist, and his team helped show how some of its elements fit together. The mechanism they studied is only one of the many processes that contribute to the much larger intestinal function of digesting food. Though his report is highly complicated, a simplified version with only a few key highlighted steps illustrates the mechanism.
Resident intestinal microorganisms work on some food types to break down complex carbohydrates into short-chain fatty acids. When intestinal epithelial cells detect short-chain fatty acids in the gut lumen, they absorb them for bodily nourishment and cell fuel. Additionally, their presence in the cell is specified by internal programming to be a signal to modify gene regulation. Long strands of DNA are kept disentangled by being wrapped around thousands of molecular “spools” called histones. There are many methods by which gene expression is regulated—one intriguing way is placing molecular markers called “crotonylations” on the histones. Crotonylations are a recent discovery. These markers are “fundamental regulators of gene expression and are tightly controlled by enzymes that respond to metabolic precursors”2 like short-chain fatty acids. One important discovery by the Varga-Weisz team was demonstrating that short-chain fatty acids increase the number of crotonylations by suppressing a protein called HDAC2.
The placement of molecules annotated on the DNA or histones as regulatory instructions are collectively called “epigenetic markers”—markers outside of the genetic code itself. In this case, specific gene products are produced corresponding to the presence of short-chain fatty acids not by altering the sequence of DNA per se, but by “turning the genes on and off,” so-to-speak, by these epigenetic controls.
Adaptation occurs internally because of superior design, not externally as a result of natural selection. ![]()
Learning about this control mechanism is fascinating, but the original paper and the news report surrounding it are also instructive to readers in their approach to understanding scientific information in two additional ways.
First, ICR recently encouraged readers to use a design-based, organism-focused approach to analyze biological systems and look for the key elements of self-adjusting systems like sensors, logic mechanisms, and output responses that enable creatures to tightly track environmental changes.3 This case illustrates how organisms use mechanisms in the digestive process to track even dietary changes.
Second, ICR urged readers to reinterpret causes reported as environmental in research papers as having engineering-based causality instead. If readers adopt this type of causal explanation, which includes observable findings and omits mystical selection events or personifications of nature “working” on creatures, then they will be attuned to spot irrational expressions of external conditions somehow “reaching” inside organisms and shaping their responses…which is common in evolutionary literature.4
A case-in-point is the headline from Babraham Institute News about Varga-Weisz’s research stating, “How good bacteria control your genes: Chemical signals from gut bacteria influence gene regulation in the gut lining.”5 It adds that researchers “have discovered a way that good bacteria in the gut can control genes in our cells…[by] chemical messages from bacteria…[to] the human genome” and that “these molecules, called short chain fatty acids, can move from the bacteria and into our own cells. Inside our cells, they can trigger processes that change gene activity and that ultimately affect how our cells behave.”
Varga-Weisz’s study showed that the entire epigenetic mechanism was internal to the cell. Short-chain fatty acids, though produced by microorganisms depending on the type of food in the gut, were only a variable that was either present or not. Since cells have membranes which tightly regulate what gets into them, then short-chain fatty acids do not just mystically move into a cell. “Good” or “bad” microorganisms don’t actively send “signals” or “messages.” Programming alone within cells determines what is or isn’t a “signal” and then that programming “triggers” the cell’s own responses. Since evolutionists believe, starting with Darwin, that active environments mold relatively passive organisms, they see the operation of biological systems backwards to how they actually operate and they include misleading, mystical language to that effect in their explanations.
This research is an example of the creative genius of the Lord Jesus who designed creatures as active, problem-solving entities which continuously track environmental changes as they solve challenges/fill new niches. ![]()
Varga-Weisz’s research is an example of the creative genius of the Lord Jesus who designed creatures as active, problem-solving entities which continuously track environmental changes as they solve challenges and fill new niches. In addition, since two independent entities like a human and a microorganism cannot work together unless connected by a specific interface system, the very interface enabling their essential symbiotic relationship is a testament to His thorough understanding of all living things as well.6
References
- Fellows, R. et al. 2018. Microbiota derived short chain fatty acids promote histone crotonylation in the colon through histone deacetylases. Nature Communications. 9 (1): 1-15.
- Fellows, 2.
- Guliuzza, R. J. 2017. Engineered Adaptability: Arriving at a Design-Based Framework for Adaptability. Acts & Facts. 46 (8): 17-19.
- Guliuzza, R. J. 2017. Engineered Adaptability: Engineering Causality Is the Answer to Darwinian Externalism. Acts & Facts. 46 (10): 17-19.
- How good bacteria control your genes. Babraham Institute News. Posted on babraham.ac.uk January 9, 2018, accessed January 9, 2018.
- Guliuzza, R. J. and F. Sherwin, 2016. Design Analysis Suggests That Our “Immune” System Is Better Understood as a Microbe Interface System. Creation Research Society Quarterly. 53 (2): 27-43.
*Randy Guliuzza is ICR’s National Representative. He earned his M.D. from the University of Minnesota, his Master of Public Health from Harvard University, and served in the U.S. Air Force as 28th Bomb Wing Flight Surgeon and Chief of Aerospace Medicine. He is also a registered Professional Engineer.
Article posted on January 25, 2018.