The concept of what comprises a gene and how it works has changed markedly since the beginning of the modern genomics era about 35 years ago when the first viral gene was sequenced.1 Since then, entire microbial, plant, and animal genomes have been sequenced.
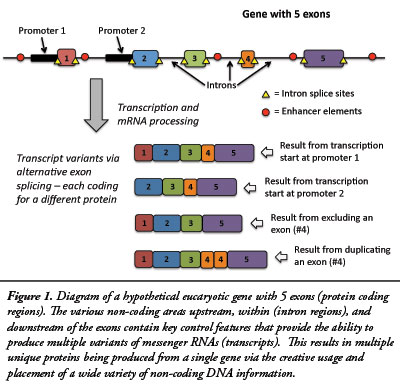
When research into gene function began, it was widely assumed that a one-to-one relationship existed between genes and their RNA and protein products. However, genome sequencing projects soon revealed that the large number of RNAs and corresponding proteins being discovered were hundreds of times more numerous than the number of genes found in the DNA sequence. We now know that this is due to the many complex mechanisms associated with gene function. In plants and animals, a gene typically produces a messenger RNA (transcript) from multiple segments of DNA in a gene region. These coding segments are called exons, while the non-coding segments (introns) are spliced out in the processing of RNA. A single gene region can produce a variety of transcripts by adding, multiplying, or eliminating exons in a process called alternative splicing (see Figure 1). For example, three neurexin genes in humans can produce over 3,000 different transcripts.2
 This author is currently summarizing key points from secular research in the area of gene function to produce a literature review for journal publication that demonstrates the irreducible complexity of gene function. This project will show that concepts of genome evolution are incredibly oversimplified, disregarding the immense levels of functional complexity unveiled by just a few decades of genomics research.
This author is currently summarizing key points from secular research in the area of gene function to produce a literature review for journal publication that demonstrates the irreducible complexity of gene function. This project will show that concepts of genome evolution are incredibly oversimplified, disregarding the immense levels of functional complexity unveiled by just a few decades of genomics research.
In brief, it is now known that gene function involves: 1) diverse regulatory DNA sequences functioning as control features located throughout gene regions, 2) complex interconnections between genes and gene networks, 3) dynamic regulation of three-dimensional chromosome architecture, 4) the interplay of DNA chemistries and conformational features, 5) cell tissue type and physiological state, and 6) the effects of DNA sequence variation within gene pools. Even these categories can be further broken down into sub-fields of study.
Scientists have attempted to deduce a predictive splicing code for many genes.3,4 This effort has been complicated by the alternative splicing between genes located on completely different chromosomes.4 For this to occur, genes in different regions of the genome are dynamically positioned within close physical proximity of each other and transcribed in highly complex gene factory zones.3 All six of the broad mechanism categories described above are involved at this level of gene function, providing a virtual symphony of unfathomable biological complexity.
Our ever-increasing knowledge of the intelligently designed genome is fully discrediting concepts of genome evolution via natural processes. The genome is an irreducibly complex system designed and implemented from the very beginning with specific uniqueness to each and every created kind, as indicated in the book of Genesis.
References
- See Sherwin, F. 2011. So, What Is a Gene? Acts & Facts. 40 (10): 16.
- U of T researchers crack “splicing code,” solve a mystery underlying biological complexity. University of Toronto news release, May 5, 2010.
- Barash, Y. et al. 2010. Deciphering the splicing code. Nature. 465 (7294): 53-59.
- Horiuchi, T. and T. Aigaki. 2006. Alternative trans-splicing: a novel mode of pre-mRNA processing. Biology of the Cell. 98 (2): 135-140.
* Dr. Tomkins is Research Associate at the Institute for Creation Research and received his Ph.D. in Genetics from Clemson University.
Cite this article: Tomkins, J. 2012. The Irreducibly Complex Genome: Designed from the Beginning. Acts & Facts. 41 (3): 6.




